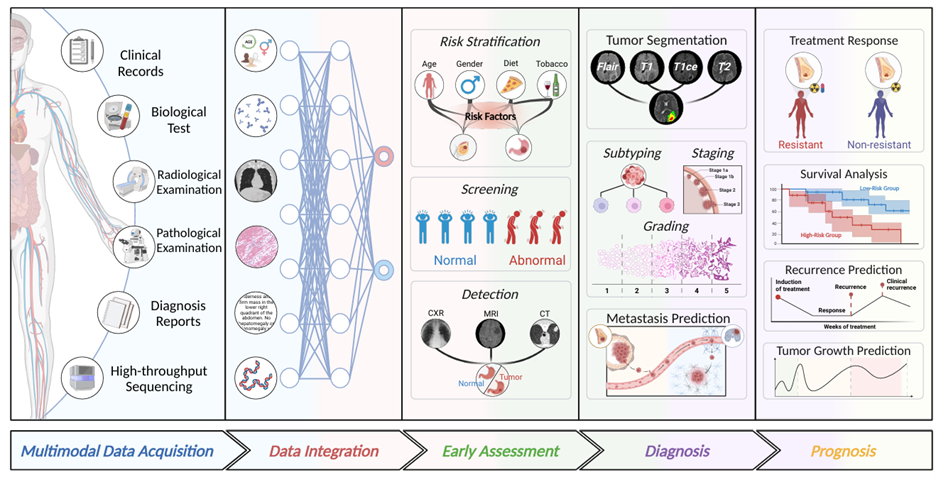
My current research focuses on multimodal AI for healthcare, aiming to advance diagnosis, prognosis, and precision medicine. I develop deep learning methods that
integrate radiomic features, tissue morphology, molecular information, and clinical records from real patient samples to
support clinical decision-making. I am particularly interested in key challenges in multimodal learning, including fusion
strategies, missing modalities, uncertainty estimation, and robustness in real-world clinical settings. More broadly,
my work explores weakly supervised and self-supervised learning under limited-label conditions, alongside core medical
image analysis tasks such as segmentation, registration, reconstruction, and AI-assisted diagnosis.